Fecal contamination of water impacts many regions of the United States and may carry risks to human health. When a water body fails to meet water quality standards for fecal bacteria, the Federal Clean Water Act requires a Total Maximum Daily Load (TMDL) analysis to establish how many bacteria are in the water, the sources of bacteria, and whether the contamination varies seasonally (See UGA CAES Publication B1242-2 – TMDLs in Georgia). The main goal is to reduce this contamination so that all waters meet regulatory standards. Each state has its own bacterial standards for water bodies, which varies depending on their use (see Extension Bulletin 1242-3. Georgia Water Quality Standards). Often the specific sources of fecal contamination in a watershed cannot be pinpointed (e.g., wildlife). Furthermore, the bacterial contribution from sediment that is resuspended during storm events is unknown. In order to adequately assess human health risks and develop watershed management plans, it is essential to identify the sources of fecal contamination.
If all the point sources of fecal contamination are acknowledged and there is still a bacterial problem, then it may be time to try additional source identification tools, such as BST. The best way to conduct BST is:
- Pinpoint the source of contamination with targeted sampling. Conduct rigorous water quality monitoring to select the specific stream reaches or tributaries that contribute to the problem. Intensive sampling, coupled with good field observations and land use information could identify the areas where fecal bacteria numbers are high. For example, if fecal coliform 1evels are high in a particular stream reach where a residential subdivision is located, it is possible to rule out livestock as the source.
- If targeted sampling yields several possible sources of fecal contamination, then BST would be applied to identify the host organism of bacteria. Most advanced techniques for identifying host organism are in the experimental stage and are often costly. Hence, it is important to pick the appropriate time and method for source identification. It is important to keep in mind that not all BST methods estimate how much each source contributes to bacterial contamination, only the different sources. In addition, there is the possibility that some sources will never be identified or may be erroneously identified.
The goal of targeted sampling is to verify which sections along a stream are potential sources of fecal contamination. The protocol, which is the same as the children’s game “hot and cold,” was developed to identify persistent sources of fecal contamination and is useful as a prelude to BST. The objective is to narrow the area where contamination is thought to originate and hopefully eliminate the need for BST. For example, if targeted sampling identifies a “hotspot” of fecal contamination that is located near a dog park and a septic field, then only those two sites and the water source must be sampled to determine the extent to which they are contributing to the fecal count. It is important to separate base flow from storm flow because fecal bacterial counts increase due to incoming runoff and sediment during storm events. This method greatly enhances the accuracy of BST. BST tests alone are typically 65-85% accurate; however, when the same tests are combined with targeted sampling, BST is even more accurate. In addition, targeted sampling is less expensive and time-consuming than BST, particularly for large areas.
It is being adopted by several Georgia Regional Development Centers. Targeted sampling does have disadvantages; it requires the agreement of property owners to cross property lines and it is labor intensive. Both drawbacks can be addressed with adequate planning and support from the community.
What is BST?
Bacterial Source Tacking (BST) is a “toolbox” of microbiological and chemical techniques to determine sources of fecal bacteria (e.g., livestock might be one source) in environmental water samples. By identifying specific sources of nonpoint pollution, BST can help enhance water quality. Also, BST data provides critical support and calibration points to improve watershed modeling. BST is considered part of Microbial Source Tracking (MST), which includes not only bacteria, but also protozoa and viruses.
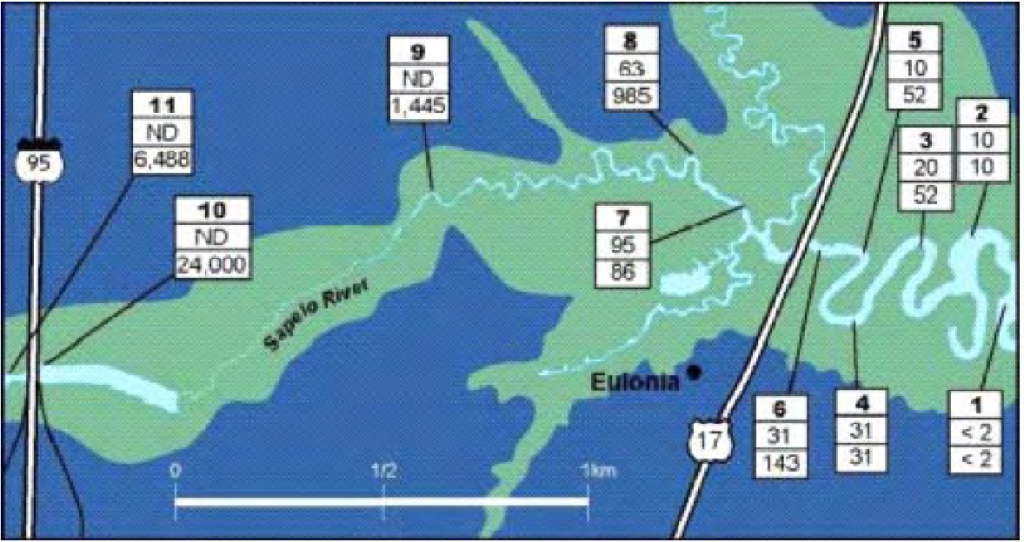
The figure below shows an example of targeted sampling with fecal enterococci counts. The top numbers are the sampling locations on the Sapelo River. The middle number represents sampling without local knowledge and the bottom number represents sampling with local knowledge (Kuntz et al., 2003).
BST Methods
BST methods can be subdivided into three basic groups: chemical, phenotypic, and genotypic. In the past, the only way to identify the host origin was to study observable characteristics of bacteria (phenotypic markers). In recent years, it has become possible to differentiate subspecies based on their DNA. This process is called genotyping.
What are fecal indicator bacteria?
Since it is almost impossible to test water for all bacteria that pose a risk to humans, the quality of water is determined by testing for the presence of indicator bacteria. The most common indicator bacteria are fecal coliforms, which are found in the intestines of warm-blooded animals and are excreted with human and animal waste. Escherichia coli (E. coli), a member of the fecal coliforms, is often used as an indicator. The presence of fecal coliforms in water bodies indicates that the water has been contaminated with the feces from of humans or other animals. Another group of fecal indicator bacteria are fecal enterococci. Several marine and freshwater studies indicate they are the best bacterial water quality indicator for brackish and marine waters.
The presence of fecal contamination is an indicator that potential health risks exist for individuals exposed to the water. The U.S. Environmental Protection Agency and other federal agencies have developed standards to determine when a water body has been contaminated. Unfortunately, these standards only quantify bacteria and do not identify their origin. Knowing the host origin of fecal coliforms is important because resources can then be directed to minimize bacterial load whenever control is possible.
Chemical methods
Chemical methods detect compounds linked to human wastewater. It is assumed that if these compounds are detected, there must be a human source associated with the contamination.
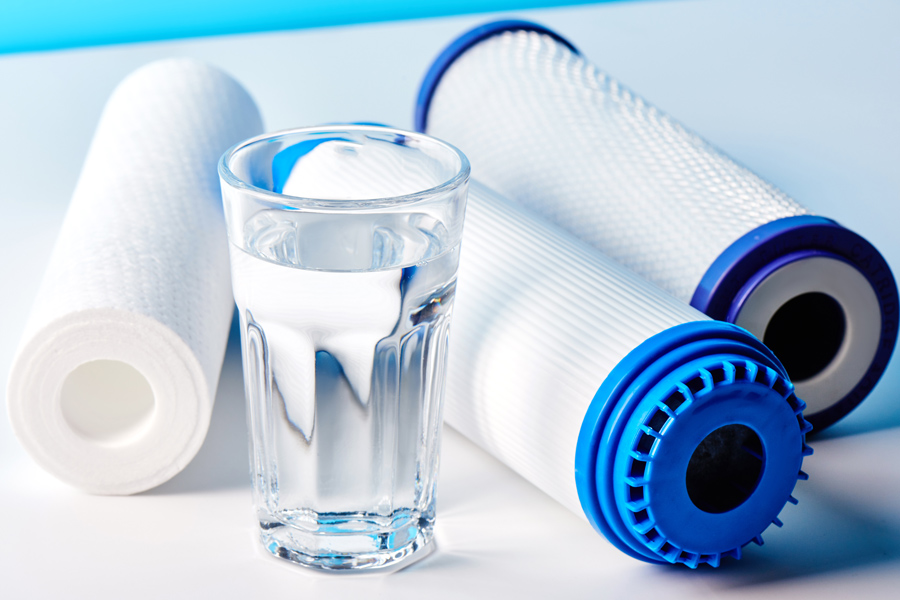
Optical brighteners are present in laundry detergents. When these compounds are exposed to ultraviolet light, they fluoresce. They persist in the environment and are measured with a fluorometer.
The problem with persistent chemicals as indicators is that they may not reflect recent pollution and they do not necessarily indicate that the water is contaminated with bacteria. Nevertheless, if optical brighteners are present, it is assumed that the stream has been affected by humans. This method may be a useful indicator of on-site septic system or gray water discharge. Although laboratory time and costs are minimal, fieldwork is time intensive.
Optical brighteners make textiles appear whiter and brighter. The tube on the left has water with optical brighteners, while the tube on the right does not.
Phenotypic methods
Phenotypic methods measure the type and quantity of substances produced by fecal bacteria. Compared with molecular methods, these methods require less training for lab personnel, have a lower cost per bacterial isolate, and can process hundreds of isolates per week. Highly contaminated sites may need several hundred isolates for the results to be representative of the fecal population in the sample. Molecular and non-molecular methods can validate each other. For example, a phenotypic method can process large number of isolates, and a molecular method can confirm the results on a few isolates.
Antibiotic Resistance Analysis (ARA) typically uses E. coli or Enterococcus species and determines their patterns of antibiotic resistance. This test relies on the premise that human fecal bacteria will have different resistance to antibiotics than bacteria in farm animals, pets, and wildlife (e.g., wildlife is expected to have little resistance to any antibiotic, since they are usually not exposed to them).
Isolates are transferred to 96-well culture plates (one isolate per well) containing a selective liquid medium. The plates are incubated, then each isolate is scored for growth or no growth. Resistance patterns will emerge with the test and sources could be differentiated.
ARA is inexpensive, fast, and can analyze large numbers of isolates. It is important to note that ARA is not able to determine where a specific source is located, only the warm-blooded animal from which it may have come.
Genotypic methods
Genotypic methods are referred to as “DNA fingerprinting” and rely on the unique genetic makeup of different strains (subspecies) of fecal bacteria. Although fecal bacteria in any two animals are genetically the same, the key to genotypic tests is finding differences in the genetic makeup against a background of similarities. The distinctions between fecal bacteria from different animals (including humans) occur because the intestinal environments and diet are not the same; hence, bacteria have evolved differences that can be related to the source. Genotypic techniques require that DNA be carefully extracted, purified, and quantified.

Ribotyping characterizes a small, specific portion of the bacterial DNA sequence. It analyzes the specific DNA sequence that codes for the production of ribosomal RNA (ribonucleic acid). Bacterial DNA is digested with a special enzyme, put in a gel, and separated. The DNA is transferred to a membrane and the membrane exposed to labeled DNA.
The DNA from E. coli will produce specific patterns that uniquely identify the bacterial strain. This method is effective in discriminating between human and non-human sources.
In contrast to ARA, ribotyping is expensive, slow, and can analyze only small numbers of isolates quickly, but its reproducibility and ability to discriminate among closely related bacteria is superior.
Does BST Work?
At this point, no single BST method is capable of identifying specific sources in all situations. Targeted sampling as a prelude to BST is recommended. If BST is needed, then a “toolbox” approach is best. Targeted sampling often eliminates the need for BST or at least narrows the scope and number of samples that have to be taken. BST can reliably determine if fecal bacteria came from human or animal sources. If the bacteria are from animal sources, BST will differentiate between livestock and wildlife, but less reliably. It is known at this time is BST can eventually achieve distinctions between different types of livestock (e.g., cattle and horse), wildlife (e.g., deer and waterfowl), or pets (e.g., cats and dogs). Future research will likely improve the accuracy of BST methods and their cost. Until then, targeted sampling provides the ability to limit the scope and number of BST tests required.
BST and Regulatory Agencies
Because BST methods appear to provide the best available technology for determining the original origin of fecal contamination in water bodies, interest in applying these techniques has grown. EPA’s recent implementation of TMDLs and projects involving TMDLs to reduce fecal coliforms would likely benefit from the use of targeted sampling and BST.
Contacts and More Information
Your County Extension Agent
https://extension.uga.edu/county-offices.html
1-800-ASK-UGA1 (directs you to the UGA Extension office in the county where your phone is registered)
EPA TMDL Program
National Small Flows Clearinghouse
https://www.nesc.wvu.edu/about-actat/national-small-flows-clearinghouse










